მოდული:Taxobox
| Sympetrum meridionale | |||
|---|---|---|---|
 | |||
| სისტემატიკა | |||
| ქვედა სამეფო | Პირველადპირიანები | ||
| superphylum | Ecdysozoa | ||
| ტიპი | Ფეხსახსრიანები | ||
| კლასი | Მწერები | ||
| რიგი | Ნემსიყლაპიები | ||
| ოჯახი | Libellulidae | ||
| გვარი | Sympetrum | ||
| სახეობა | S. meridionale | ||
| Ლათინური სახელი - სახეობა | |||
| Sympetrum meridionale | |||
| Selys, 1841 | |||
| დაცვის სტატუსი | |||
 | |||
| |||
| Python Royal | |||
|---|---|---|---|
 | |||
| systématique | |||
| clade | Episquamata | ||
| clade | Toxicofera | ||
| clade | Ophidia | ||
| sous-ordre | Serpentes | ||
| infra-ordre | Alethinophidia | ||
| clade | Afrophidia | ||
| super-famille | Pythonoidea | ||
| famille | Pythonidae | ||
| genre | Python | ||
| espèce | P. regius | ||
| Scientific name of espèce | |||
| Python regius | |||
| Shaw, 1802 | |||
| synonyme | |||
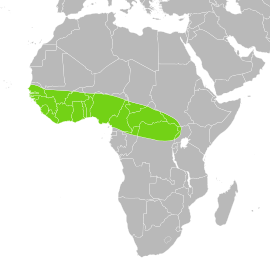
| Range map | |||
 | |||
| statut de conservation UICN | |||
 | |||
| |||
This module is still unstable. Use with your own caution and report bugs and feature requests at Module talk:Taxobox or Wikidata talk:WikiProject_Taxonomy.
Taxobox.lua is a lua module which can automatically generate taxonomy infobox and is overwritable by classic taxobox parameters like species, unranked_ordo etc.
This module infobox is designed to be a replacement of Wikipedia's Taxobox. It provides configuration options which can control hypernym paths, show or hide certain ranks, specify content language and its configs, and make a callback and pass parameters to "classic" taxobox.
The following code
{{Taxobox
| qid=Q464424
}}
creates the taxobox on the right hand side. The item to show is given with qid.
If you want to have this taxobox show up on each Wikidata taxon item: [1]
Internal რედაქტირება
Method taxobox
რედაქტირება
The taxobox method provides function runs the above example. The method itself can be invoked with
{{#invoke: Taxobox
| taxobox
| qid=Q464424
| config[count]=10
}}
The number of parent taxons to show is given by config[count].
Method callback
რედაქტირება
The callback method provides function to retrieve all internal parameters and pass them to an external template. The method can be invoked with like this
{{#invoke: Taxobox
| callback
| qid=Q464424
| template=OtherTaxobox
| config[count]=10
}}
callback accepts all arguments that taxobox accepted. It also accepts an extra argument template to specify name of the template to be expanded.
I18n რედაქტირება
Change the i18n messages in Module:I18n/taxobox. I18n also specifys some format strings which can be use to customize the infobox output of certain language.
| message | description | example(s) | |
|---|---|---|---|
rank-format
|
Format of instances of
|
, or
713623 |
. Will be passed into 2 named arguments when rendering ranks:
{link}- Wiki link to the rank
{label}- Display title of the rank
"[[{link}|{label}]]": display name and give a hyperlink to the rank, or"{label}"only display name of the rank
rank-format-<latinrank>
This argument is similar to rank-format but can be use to specify the format of certain rank. The "<latinrank>" is a latin name, can be a instance of
427626 |
, or
713623 |
. For example "rank-format-cladus" for
713623 |
. It accepts same 2 named arguments like rank-format
"": don't display anything at this rank. Can be used to hide the rank name when the taxon is a clade"<i>(clade)</i>": use italic style"[[{link}|<span style=\"color:gray\">{label}</span>]]": use link and change color
item-format-current-with-vernacular-name
Format of instances of
16521 |
or
310890 |
. current means either the item is the main taxon (specified by module argument qid), or the item and all taxa between the item and main taxon (if any) are all
310890 |
. with-vernacular-name means the item's 1843 or label exists and is different from 225.
4 named arguments will be passed on redering:
{link}- Wiki link to the taxon item
{vernacular}- The "common name" (the value of 1843 or item label) of taxon item in current language
{scientific}- Full scientific name
{scientificshort}- Short scientific name, for example "P. leo" of
140 |
Note: Don't use italic style here on scientific names. This style can be set by scientific-name-pattern, scientific-name-repl and other related messages.
"<b>{scientificshort}</b>": only display the short scientific name in bold weight"<b>{vernacular} {scientificshort}</b>": display both vernacular and short scientific name in bold weight"[[{link}|{vernacular}]]": display a hyperlink to target taxon with its vernacular name as label
item-format-current-with-vernacular-name
Format of instances of
16521 |
or
310890 |
. without-vernacular-name means either both 1843 and label (in specified language) are empty, or the vernacular name is the same as 225.
4 named arguments will be passed on rendering, and they are same to item-format-current-with-vernacular-name.
"[[{link}|{scientificshort}]]": display a hyperlink to target taxon with its short scientific name as label"[[{link}|{vernacular}]] ({scientficshort})": display both vernacular and short scientific name, also link to target taxon on from vernacular name
scientific-name-pattern, scientific-name-repl
Pattern replacement to generate full scientific name. Can be override per rank by scientific-name-pattern-<latinrank> and scientific-name-repl-<latinrank>.
Not the format of scientific-name-pattern and other *pattern* messages are Lua's patterns, which are similar to
125267 |
but not identical. Read the manual to know how to write ones.
- (pattern)
"^.+$": match all non-empty strings- (repl)
"%0": do nothing for replacement, or - (repl)
"<i>%0</i>": put the scientific name in <i> tag so it can be rendered as italic
- (repl)
short-scientific-name-pattern, short-scientific-name-repl
Pattern replacement to generate short scientific name. Can be override per rank by short-scientific-name-pattern-<latinrank> and short-scientific-name-repl-<latinrank>.
scientific-name-pattern-<latinrank>, scientific-name-repl-<latinrank>
Pattern replcacement to generate full scientific name per rank. The "<latinrank>" is a latin name, can be a instance of
427626 |
, or
713623 |
. For example "scientific-name-pattern-species" for
7432 |
.
- (
scientific-name-pattern-genus)nil: fallback toscientific-name-pattern's"^.+$"- (
scientific-name-repl-genus)"<i>%0</i>": italicize full scientific name if its rank is
- (
34740 |
short-scientific-name-pattern-<latinrank>, short-scientific-name-repl-<latinrank>
Pattern replcacement to generate short scientific name per rank. The "<latinrank>" is a latin name, can be a instance of
427626 |
, or
713623 |
. For example "short-scientific-name-pattern-species" for
7432 |
.
- (
short-scientific-name-pattern-species)"^(%w)%w+ (%w+)$": pattern to fetch the first letter of genus name (as %1) and the whole epithet (as %2) from the binomial of a species.- (
short-scientific-name-repl-species)"<i>%1. %2</i>": for example, the result for
- (
140 |
is "P. leo"
scientific-name-replaces or short-scientific-name-replaces
Pattern replacements to apply for all full (or short) scientific names, after the name has been processed by pattern-repl pair described above.
The value for each of two messages is not a string but a table. The table contains multiple pattern-repl pairs which will be applied to scientific names.
Note: Lua's table object doesn't sort, so the replacement sequence CAN NOT be guaranteed. DON'T DEPEND ON THE SEQUENCE YOU SAW!
Input Parameters რედაქტირება
The "<latinrank>" below is a latin name, can be an instance of
427626 |
, or
713623 |
. For example "display[cladus]".
Config Options რედაქტირება
config[lang]: content language (default:en).config[count]: maximum count of taxon to be recursively iterated (default: 10).config[references]: a space-separated list of item ids. The references to favor in case of alternative claims. Optional.config[usetaxa]: a space-separated list of item ids. The taxa to favor in case of alternative claims. Optional.config[link]: if the value is "sitelink" it will use local wiki site links instead of wikidata item links.config[dryun]: used forcallbackmethod. Displaying a <pre> block contains wikitext instead of expanding and rendering the template. Dryrun can be used to find parameters to be overrided.
Examples რედაქტირება
| Ლომი | |||
|---|---|---|---|
 | |||
| სისტემატიკა | |||
| grandorder | Ferae | ||
| რიგი | Მტაცებლები | ||
| ქვერიგი | Feliformia | ||
| ოჯახი | Კატისებრნი | ||
| ქვეოჯახი | Დიდი კატები | ||
| გვარი | Პანთერა | ||
| სახეობა | Ლომი | ||
| Ლათინური სახელი - სახეობა | |||
| Panthera leo | |||
| Linnaeus, 1758 | |||
| სინონიმი (ტაქსონომია) | |||
| გავრცელების არეალი | |||
 | |||
| დაცვის სტატუსი | |||
 | |||
| აუდიო | |||
| |||
{{#invoke: taxobox
| taxobox
| qid = Q140
| config[usetaxa] = Q27379
|config[count] = 7
}}
Display Options რედაქტირება
display[<latinrank>]: if the value is "n", "no", "false" or "hide", the specified rank (in latin name or QID) will be hide. Otherwise the rank will display.
For example, to hide all clades:
| Ლომი | |||
|---|---|---|---|
 | |||
| სისტემატიკა | |||
| ზერიგი | Laurasiatheria | ||
| grandorder | Ferae | ||
| რიგი | Მტაცებლები | ||
| ქვერიგი | Feliformia | ||
| ოჯახი | Კატისებრნი | ||
| ქვეოჯახი | Დიდი კატები | ||
| გვარი | Პანთერა | ||
| სახეობა | Ლომი | ||
| Ლათინური სახელი - სახეობა | |||
| Panthera leo | |||
| Linnaeus, 1758 | |||
| სინონიმი (ტაქსონომია) | |||
| გავრცელების არეალი | |||
 | |||
| დაცვის სტატუსი | |||
 | |||
| აუდიო | |||
| |||
{{#invoke:taxobox
|taxobox
|qid=Q140
|display[cladus]=hide
}}
Classic Parameters რედაქტირება
<latinrank>: The classic taxon parameters used by most
52496 |
. All of them can be overrided manually.
<latinrank>_authority: The classic taxon authority parameters used by most
52496 |
. All of them can be overrided manually.
unranked_<latinrank>: The classic unranked parameters used by most
52496 |
. All of them can be overrided manually.
If there are more than one clades between two taxon ranks, you can override them by appending [<number>] index to the unranked_<latinrank> parameter.
For example, there are 3 clades between rank genus and rank species. You can override them like this:
{{#invoke:taxobox
|taxobox
|...
|unranked_species[3] = Cladus closest to genus rank
|unranked_species[2] = The middle clade
|unranked_species[1] = Cladus closest to species rank
|...
}}
Output Parameters რედაქტირება
The best way to see all output parameters is to use the config[dryrun] parameter:
{{#invoke:taxobox
|callback
|qid=Q140
|config[dryrun]=yes
}}
The result:
{{Taxobox
|audio = Lion raring-sound1TamilNadu178.ogg
|code = 13011
|color = pink
|config[dryrun] = yes
|image = 002 The lion king Snyggve in the Serengeti National Park Photo by Giles Laurent.jpg
|iucn_status[id] = 278113
|iucn_status[image] = Status iucn3.1 VU.svg
|iucn_status[label] = მოწყვლადი სახეობები
|iucn_status[references] = Q115962546
|name = Ლომი
|qid = Q140
|range_map = Lion distribution.png
|rank[1][id] = 5868144
|rank[1][is_extinct] = no
|rank[1][is_monotypic] = no
|rank[1][is_subject] = no
|rank[1][latin] = superordo
|rank[1][link] = Q27379
|rank[1][raw_scientific] = Laurasiatheria
|rank[1][references] = Q28206255
|rank[1][scientific] = Laurasiatheria
|rank[1][taxon] = [[d:Q27379|Laurasiatheria]]
|rank[2][id] = 713623
|rank[2][is_extinct] = no
|rank[2][is_monotypic] = no
|rank[2][is_subject] = no
|rank[2][latin] = cladus
|rank[2][link] = Q7439311
|rank[2][raw_scientific] = Scrotifera
|rank[2][references] = Q30699754
|rank[2][scientific] = Scrotifera
|rank[2][taxon] = [[d:Q7439311|Scrotifera]]
|rank[3][id] = 713623
|rank[3][is_extinct] = no
|rank[3][is_monotypic] = no
|rank[3][is_subject] = no
|rank[3][latin] = cladus
|rank[3][link] = Q5444079
|rank[3][raw_scientific] = Fereuungulata
|rank[3][references] =
|rank[3][scientific] = Fereuungulata
|rank[3][taxon] = [[d:Q5444079|Fereuungulata]]
|rank[4][id] = 6462265
|rank[4][is_extinct] = no
|rank[4][is_monotypic] = no
|rank[4][is_subject] = no
|rank[4][latin] = grandordo
|rank[4][link] = Q20868
|rank[4][raw_scientific] = Ferae
|rank[4][references] = Q21608408
|rank[4][scientific] = Ferae
|rank[4][taxon] = [[d:Q20868|Ferae]]
|rank[5][id] = 36602
|rank[5][is_extinct] = no
|rank[5][is_monotypic] = no
|rank[5][is_subject] = no
|rank[5][latin] = ordo
|rank[5][link] = Q25306
|rank[5][raw_scientific] = Carnivora
|rank[5][references] = Q82575 Q1538807 Q19604469 Q21682704 Q19302303
|rank[5][scientific] = Carnivora
|rank[5][taxon] = [[მტაცებლები|Მტაცებლები]]
|rank[5][vernacular] = Მტაცებლები
|rank[6][id] = 5867959
|rank[6][is_extinct] = no
|rank[6][is_monotypic] = no
|rank[6][is_subject] = no
|rank[6][latin] = subordo
|rank[6][link] = Q27070
|rank[6][raw_scientific] = Feliformia
|rank[6][references] = Q19302303 Q19604469 Q1538807 Q82575
|rank[6][scientific] = Feliformia
|rank[6][taxon] = [[d:Q27070|Feliformia]]
|rank[7][id] = 35409
|rank[7][is_extinct] = no
|rank[7][is_monotypic] = no
|rank[7][is_subject] = no
|rank[7][latin] = familia
|rank[7][link] = Q25265
|rank[7][raw_scientific] = Felidae
|rank[7][references] = Q82575 Q1538807 Q19604469 Q19771288 Q19302303
|rank[7][scientific] = Felidae
|rank[7][taxon] = [[კატისებრნი|Კატისებრნი]]
|rank[7][vernacular] = Კატისებრნი
|rank[8][id] = 164280
|rank[8][is_extinct] = no
|rank[8][is_monotypic] = no
|rank[8][is_subject] = no
|rank[8][latin] = subfamilia
|rank[8][link] = Q230177
|rank[8][raw_scientific] = Pantherinae
|rank[8][references] = Q82575 Q19604469 Q1538807 Q56211181
|rank[8][scientific] = Pantherinae
|rank[8][taxon] = [[დიდი კატები|Დიდი კატები]]
|rank[8][vernacular] = Დიდი კატები
|rank[9][id] = 34740
|rank[9][is_extinct] = no
|rank[9][is_monotypic] = no
|rank[9][is_subject] = no
|rank[9][latin] = genus
|rank[9][link] = Q127960
|rank[9][raw_scientific] = Panthera
|rank[9][references] = Q82575 Q1538807 Q19604469 Q43385197 Q77980496
|rank[9][scientific] = <i>Panthera</i>
|rank[9][taxon] = [[პანთერა|Პანთერა]]
|rank[9][vernacular] = Პანთერა
|rank[10][authority] = [[კარლ ლინე|Linnaeus]], 1758
|rank[10][id] = 7432
|rank[10][is_extinct] = no
|rank[10][is_monotypic] = no
|rank[10][is_subject] = yes
|rank[10][latin] = species
|rank[10][link] = Q140
|rank[10][raw_scientific] = Panthera leo
|rank[10][references] = Q1538807
|rank[10][scientific] = <i>Panthera leo</i>
|rank[10][taxon] = <b>Ლომი</b>
|rank[10][vernacular] = Ლომი
|rank[references] = Q30135917 Q1538807 Q17073815 Q22814967 Q30136284 Q22826076 Q30136117 Q30135809 Q19302303 Q33883775 Q50754531 Q28191107 Q796451 Q82575
|rank[size] = 10
|synonym[1][author] = [[კარლ ლინე|Linnaeus]], 1758
|synonym[1][link] = d:Q15294488
|synonym[1][name] = Felis leo
|synonym[size] = 1
|taxon = Panthera leo
}}
All output parameters can be overrided by specifying same name input parameters. For example this will replace
34740 |
to "Foo" and
2455704 |
"Bar":
| Ლომი | |||
|---|---|---|---|
 | |||
| სისტემატიკა | |||
| ზერიგი | Laurasiatheria | ||
| კლადი | Scrotifera | ||
| კლადი | Fereuungulata | ||
| grandorder | Ferae | ||
| რიგი | Მტაცებლები | ||
| ქვერიგი | Feliformia | ||
| ოჯახი | Კატისებრნი | ||
| ქვეოჯახი | Bar | ||
| გვარი | Foo | ||
| სახეობა | Ლომი | ||
| Ლათინური სახელი - სახეობა | |||
| Panthera leo | |||
| Linnaeus, 1758 | |||
| სინონიმი (ტაქსონომია) | |||
| გავრცელების არეალი | |||
 | |||
| დაცვის სტატუსი | |||
 | |||
| აუდიო | |||
| |||
{{#invoke:taxobox
|taxobox
|qid=Q140
|subfamilia=<strong style="color: green">Bar</strong>
|rank[9][taxon]=<strong style="color: red">Foo</strong>
}}
Supported properties რედაქტირება
The taxobox currently supports:
- 18 for taxon images and red list status
- 31: for
310890 |
and
14594740 |
- 105
- 141 also mark the taxon extinct if the value is
237350 |
- 171
- 181
- 225
- 248 for references
- 405
- 428
- 523
- 524 also mark the taxon extinct if present
- 566
- 574
- 582 mark the taxon extinct if present
- 697
- 428 if ICNafp applies, otherwise:
- 835, otherwise the last name of English language (en) is shown.
- 944 for authority string format and color.
- 1843, common name of a language to override the item label of the language
Wikipedia use რედაქტირება
This module is designed to be a replacement for Wikipedia taxoboxes. However, it is still unstable and need plenty extra template works to allow a classic
52496 |
to accept the new callback parameters. Suggestion and bug reports are welcome at d:Module talk:Taxobox or d:Wikidata talk:WikiProject_Taxonomy.
სქოლიო რედაქტირება
-- vim: set noexpandtab ft=lua ts=4 sw=4:
local ENABLE_DEBUG = true
--local Cite = require('Module:Cite')
local fb = require('Module:Fallback')
local p = {} -- module exports
local L = {} -- alias to local functions
-- (so it can be iterated by p in debug mode)
local _linkconfig -- use links from content language Wikipedia
-- or from Wikidata, default to Wikidata
local _contentlang
local usereferences -- array of references to be prefered in the given order
local usetaxa -- array of taxa to be preferred in the given order
local hideranks -- array of ranks to be show or hide from display
local code = false
local subcode = false
local visited = {}
-- biological nomenclatures
NOMENCLATURE_ICZN = 13011 -- Zoo
NOMENCLATURE_ICNafp = 693148 -- Algae, Fungi and Plants
NOMENCLATURE_ICNCP = 764 -- Cultivated Plants
NOMENCLATURE_ICNP = 743780 -- Prokaryota/Bacteria
NOMENCLATURE_ICVCN = 14920640 -- Viruses
-- look for taxons for color selection
ARCHAEA = 10872 -- ICNP
FUNGI = 764 -- ICNafp
SAR = 137323
HAROSA = 18397957 -- SAR
CHROMALVEOLATA = 477950 -- SAR
CHROMISTA = 862296 -- SAR
RHIZARIA = 855740 -- SAR
AMOEBOZOA = 473809 -- Eukaryota
EXCAVATA = 691551 -- Eukaryota
EUKARYOTA = 19088
-- background colors for each code
local colors = {
[false] = '#d3d3d3',
[NOMENCLATURE_ICZN] = 'pink',
[NOMENCLATURE_ICNafp] = 'lightgreen',
[NOMENCLATURE_ICNCP] = 'lightgreen',
[NOMENCLATURE_ICNP] = 'lightgrey',
[NOMENCLATURE_ICVCN] = 'violet',
[ARCHAEA] = '#c3f5fa',
[FUNGI] = 'lightblue',
[SAR] = '#c8fa50',
[HAROSA] = '#c8fa50',
[CHROMALVEOLATA] = '#c8fa50',
[CHROMISTA] = '#c8fa50',
[RHIZARIA] = '#c8fa50',
[AMOEBOZOA] = '#f5d7ff',
[EXCAVATA] = '#f5d7ff',
[EUKARYOTA] = '#f5d7ff',
}
local virusgroups = {
[2901600] = {group = 'I', shortlabel = 'dsDNA'},
[9094469] = {group = 'II', shortlabel = 'ssDNA'},
[3307900] = {group = 'III', shortlabel = 'dsRNA'},
[9094478] = {group = 'IV', shortlabel = 'ssRNA(+)'},
[9285327] = {group = 'V', shortlabel = 'ssRNA(-)'},
[9094482] = {group = 'VI', shortlabel = 'ssRNA-RT'},
[3754200] = {group = 'VII', shortlabel = 'dsDNA-RT'},
[44209729] = {group = nil, shortlabel = 'ssDNA(-)'},
[44209788] = {group = nil, shortlabel = 'ssDNA(+)'},
[44209909] = {group = nil, shortlabel = 'ssDNA(+/-)'},
[209917] = {group = nil, shortlabel = 'Viroid'},
[44209519] = {group = nil, shortlabel = 'ssRNA(+/-)'},
[45181439] = {group = nil, shortlabel = 'ssRNA'},
}
local i18nmessages = require("Module:I18n/taxobox")
-- readable taxon properties
local P_IMAGE = "P18"
local P_INSTANCE_OF = "P31"
local P_TAXON_RANK = "P105"
local P_IUCN_STATUS = "P141"
local P_TAXON_PARENT = "P171"
local P_SPREAD_MAP = "P181"
local P_TAXON_NAME = "P225"
local P_STATED_IN = "P248"
local P_AUTHOR = "P405"
local P_AUTHOR_ABBR_IPNI = "P428"
local P_ERA_START = "P523"
local P_ERA_END = "P524"
local P_BASIONYM = "P566"
local P_SYNONYM = "P1420"
local P_REPLACED_SYNONYM ="P694"
local P_ORIGINAL_COMBINATION = "P1403"
local P_TAXON_YEAR = "P574"
local P_START_TIME = "P580"
local P_END_TIME = "P582"
local P_EX_AUTHOR = "P697"
local P_AUTHOR_ABBR_ZOOLOGY = "P835"
local P_NOMENCLATURE_CODE = "P944"
local P_COMMON_NAME = "P1843"
local P_AUDIO = "P51"
local P_INCERTAE_SEDIS = "P678"
local P_TAXONOMIC_TYPE = "P427"
local P_VIRUS_GENOME = "P4628"
local P_SUBJECT_ROLE = "P2868"
local P_OF = "P642"
local COMMONS_CATEGORY="P373"
-- readable item
local CLADE = 713623
local GENUS = 34740
local SUBGENUS = 3238261
local ZOOSECTIO = 10861426
local ZOOSUBSECTIO = 10861375
local RED_DATA_LIST = 32059
local MONOTYPIC_TAXON = 310890
local GEOLOGICAL_ERA = 630830
local SYSTEMATICS = 3516404
local RECOMBINATION = 14594740
local EXTINCT = 237350
local INCERTAE_SEDIS = 235536
local SYNONYM_TAXON = 1040689
local TYPE_GENUS = 842832
local TYPE_SPECIES = 252730
local VIRUS_CLASSIFICATION = 478216
local PROTONYM = 14192851
local BASIONYM = 810198
local FOSSIL_TAXON = 23038290
local ET_AL = 311624
local function capitalize(text)
return mw.ustring.gsub(text, "^%l", mw.ustring.upper)
end
local function mergeTable(a, b)
for _, value in ipairs(b) do
a[#a + 1] = value
end
return a
end
L.mergeTable = mergeTable
-- credit to http://lua-users.org/wiki/StringInterpolation
local function namedStringFormat(str, vars)
-- Allow replace_vars{str, vars} syntax as well as
-- replace_vars(str, {vars})
if not vars then
vars = str
str = vars[1]
end
return (string.gsub(str, "({([^}]+)})",
function(whole,i)
return vars[i] or whole
end))
end
L.namedStringFormat = namedStringFormat
local function setLang(contentlang)
_contentlang = contentlang or mw.language.getContentLanguage():getCode()
end
L.setLang = setLang
local function getLang()
return _contentlang
end
L.getLang = getLang
local function i18n(str)
local message = i18nmessages[str]
if type(message) == 'string' then
return message
end
return fb._langSwitch(message, getLang())
end
L.i18n = i18n
-- parse item-ids like argument (like config[references]) which is a space
-- separated list of item numbers like "Q1 Q2 Q3"
local function parseItemIds(itemids)
local items = {}
local priority = 0
if itemids then
for word in string.gmatch(itemids, "%w+") do
priority = priority + 1
item = "Q" .. tonumber(string.sub(word, 2))
items[item] = priority
end
end
items.size = priority
return items
end
L.parseItemIds = parseItemIds
-- parse config arguments passed by #invode:taxobox. below are all we
-- support currently:
-- - config[lang]: set content language (default: en)
-- - config[count]: maximum count of taxon to be recursively iterated
-- - config[references]: references to be preffered in the given order
-- - config[dryrun]: generate <pre> block instead of expanding template
-- - config[link]: local or wikidata
local function parseConfig(args)
setLang(args["config[lang]"])
local count = tonumber(args["config[count]"]) or 10
if count > 25 then
-- count = 25 is roughly about 100 expensive parser function calls
error(i18n("taxon-count-too-high"))
end
usereferences = parseItemIds(args["config[references]"])
usetaxa = parseItemIds(args["config[usetaxa]"])
hideranks = {}
local displaypattern = "^display%[([^]]+)%]$"
local qidpattern = "^Q?(%d+)$"
for k, v in pairs(args) do
v = mw.ustring.lower(v)
if string.match(k, displaypattern) then
k = string.gsub(k, displaypattern, "%1")
if string.match(k, qidpattern) then
k = string.gsub(k, qidpattern, "%1")
k = tonumber(k)
end
-- TODO: i18n?
if ({n=true, no=true, ["false"]=true, hide=true})[v] then
hideranks[k] = true
end
end
end
_linkconfig = string.match(mw.site.server, "wikidata") or "sitelink"
if args["config[link]"] and
mw.ustring.lower(args["config[link]"]) == "sitelink" then
_linkconfig = "sitelink"
end
return {
["count"] = count,
["lang"] = lang,
["dryrun"] = args["config[dryrun]"],
["link"] = _linkconfig
}
end
-- Adopted from source: c:Module:Wikidata_label
-- the label of the item if present in the specified language or 'no label'
local function getLabel(item, lang)
local entity, label, language
lang = lang or getLang()
if type(item) == "number" then
item = "Q" .. item
end
if type(item) ~= 'string' then -- "item" is not a q-code
entity = item -- "item" must be the entity
item = entity.id -- look-up q-code
end
local userLang = mw.getCurrentFrame():callParserFunction( "int", "lang" )
-- get label (visible part of the link)
if (userLang == lang) and (not entity) then -- call if requesting label in user's language, but skip if we already have entity
label, language = mw.wikibase.getLabelWithLang(item) -- prefered way of calling that, as not needed to load the entire entity
end
-- hard way to get label by querying all the languages on the langList
if not label then -- used if requesting label in language different than user's, or if we already have entity
entity = entity or mw.wikibase.getEntity(item) -- load entity if we do not have it yet
label = entity:getLabel(lang) or i18n('no-label')
end
return label
end
L.getLabel = getLabel
local function getLink(id, label, format, named, ucfirst)
if type(id) == "number" then
id = "Q" .. id
end
local link = id
if _linkconfig == "sitelink" then
link = mw.wikibase.sitelink(id) or "d:" .. id
end
label = label or getLabel(id)
label = ucfirst and label or label
format = format or "[[%s|%s]]"
if named then
return namedStringFormat{format, link=link, label=label}
else
return string.format(format, link, label)
end
end
L.getLink = getLink
local function referenceTargetIds(references, property)
local ids = {}
if references then
for _, ref in pairs(references) do
for _, snak in pairs(ref.snaks[property] or {}) do
if snak.datavalue and snak.datavalue.value then
ids[tostring('Q' .. snak.datavalue.value['numeric-id'])] = true
mw.log('string ref', snak.datavalue.value['numeric-id'])
end
end
end
end
return ids
end
L.referenceTargetIds = referenceTargetIds
-- Collect all claims of the given property of the item
-- Returns all claims, their references and qualifiers in tables combined by the claims' rank.
-- result.preferred[target id of claim] = [target id of P248 reference]
-- use only if the data type of the property is item
local function targetIds(item, property)
local claims = {preferred = {}, normal = {}, deprecated = {}}
if item and item.claims and item.claims[property] then
for _,claim in pairs(item.claims[property]) do
local valueid = claim.mainsnak.datavalue and claim.mainsnak.datavalue.value
and claim.mainsnak.datavalue.value['numeric-id']
or 'novalue'
local refids = referenceTargetIds(claim.references, P_STATED_IN)
claims[claim.rank][valueid] = {refids = refids or true, qualifiers = claim.qualifiers}
end
end
return claims
end
L.targetIds = targetIds
-- Gives the first highest ranked claim and its references.
-- use only if the data type of the property is item
local function targetId(item, property)
local claims = targetIds(item, property)
if next(claims.preferred) then
return claims.preferred
end
if next(claims.normal) then
return claims.normal
end
return claims.deprecated
end
L.targetId = targetId
-- Collect all claims of the given property of the item
-- Returns a triple of claims, their qualifiers, and their references in tables combined by the claims' rank.
-- Use only if the data type of the property is string
local function targetStrs(item, property)
choosenclaim = {preferred = {}, normal = {}, deprecated = {}}
choosenqualifiers = {preferred = {}, normal = {}, deprecated = {}}
choosenreferences = {preferred = {}, normal = {}, deprecated = {}}
if item and item.claims and item.claims[property] then
for _,claim in pairs(item.claims[property]) do
if claim.mainsnak and claim.mainsnak.datavalue then
index = #choosenclaim[claim.rank] + 1
mw.log(index, claim.mainsnak.datavalue.value)
local refids = referenceTargetIds(claim.references, P_STATED_IN)
if claim.mainsnak.datatype == 'monolingualtext' then
choosenclaim[claim.rank][index] = tostring(claim.mainsnak.datavalue.value)
else
choosenclaim[claim.rank][index] = tostring(claim.mainsnak.datavalue.value)
end
choosenqualifiers[claim.rank][index] = claim.qualifiers
choosenreferences[claim.rank][index] = refids
end
end
end
return choosenclaim, choosenqualifiers, choosenreferences
end
L.targetStrs = targetStrs
-- Gives the first highest ranked claim and its qualifiers and references.
-- Use only if the data type of the property is string
local function targetStr(item, property)
choosenclaim, choosenqualifiers, choosenreferences = targetStrs(item, property)
for _, priority in pairs({"preferred", "normal", "deprecated"}) do
local index = next(choosenclaim[priority])
if index then
return choosenclaim[priority][index],
choosenqualifiers[priority][index],
choosenreferences[priority][index]
end
end
return
end
L.targetStr = targetStr
-- helper function to merge all claims, regardless of rank
local function mergeClaims(claims, qualifiers, references)
local c = {}
local q = {}
local r = {}
for _, priority in pairs({"preferred", "normal", "deprecated"}) do
mergeTable(c, claims[priority] or {})
mergeTable(q, qualifiers[priority] or {})
mergeTable(r, references[priority] or {})
end
return c, q, r
end
L.mergeClaims = mergeClaims
local function targetValue(item, property)
if item and item.claims and item.claims[property] then
for _,claim in pairs(item.claims[property]) do
if claim.mainsnak and claim.mainsnak.datavalue then
return claim.mainsnak.datavalue.value
end
end
end
end
L.targetValue = targetValue
-- same as targetId but for qualifiers
-- TODO merge
local function qualifierTargetId(qualifiers, property)
local claims = {}
if qualifiers and qualifiers[property] then
for _,claim in pairs(qualifiers[property]) do
local valueid = claim.datavalue.value['numeric-id']
table.insert(claims, valueid)
end
end
return claims
end
L.qualifierTargetId = qualifierTargetId
-- same as targetValue but for qualifiers
local function qualifierTargetValue(qualifiers, property)
local claims = {}
if qualifiers and qualifiers[property] then
for _,claim in pairs(qualifiers[property]) do
if claim.datavalue then
return claim.datavalue.value
end
end
end
end
L.qualifierTargetValue = qualifierTargetValue
-- takes a list of item ids (the values of the given table) and creates wikilinks based on their labels
local function createLinks(list, authorAbbreviation)
local authors = {}
for _,authorid in pairs(list) do
if authorid then
local author = mw.wikibase.getEntity('Q' .. authorid)
if author then
local label
if authorAbbreviation then
if code == NOMENCLATURE_ICNafp then
-- get author abbrieviation per IPNI set
if targetStr(author, P_AUTHOR_ABBR_IPNI) then
label = targetStr(author, P_AUTHOR_ABBR_IPNI)
end
elseif targetStr(author, P_AUTHOR_ABBR_ZOOLOGY) then
-- get zoologist author citation set
label = targetStr(author, P_AUTHOR_ABBR_ZOOLOGY)
end
if not label then
-- use the "last" name if no abbreviation found
-- also don't use the translated name
label = getLabel(author, "ka")
if label ~= i18n('no-label') then
_, _, label = mw.ustring.find(label, "(%w+)$")
end
end
end
table.insert(authors, getLink(authorid, label))
end
end
end
return authors
end
L.createLinks = createLinks
local function vernacularName(item)
local vernacularname
-- select vernacular name for current language
if item.claims and item.claims[P_COMMON_NAME] then
for _, claim in pairs(item.claims[P_COMMON_NAME]) do
if claim.mainsnak and claim.mainsnak.datavalue and
claim.mainsnak.datavalue.type == "monolingualtext" and
claim.mainsnak.datavalue.value.language == getLang() then
vernacularname = claim.mainsnak.datavalue.value.text
break
end
end
if vernacularname == '' then
vernacularname = nil
end
end
if not vernacularname then
-- test if item label is not one of the scientific names
vernacularname = getLabel(item)
scnames = mergeClaims(targetStrs(item, P_TAXON_NAME))
for _, n in pairs(scnames) do
if vernacularname == n then
return
end
end
end
if vernacularname == i18n("no-label") then
return
end
return capitalize(vernacularname)
end
L.vernacularName = vernacularName
local function authorString(item, namequalifiers, pid)
pid = pid or P_AUTHOR -- set default property
local concatstr = ', '
local authorids = qualifierTargetId(namequalifiers, pid) -- get qualifiers
if not next(authorids) then -- no qualifiers found, check properties
local authorset = targetId(item, pid)
local authors = {}
if authorset then -- create list from set
authorids = {}
for author,_ in pairs(authorset) do
table.insert(authorids, author)
end
end
end
local authors = createLinks(authorids, true)
if next(authors) then
local authorstr = ''
local comma = false
for i = #authors, 1, -1 do
local sep = ''
if i > 1 then
sep = ' '
if authorids[i] ~= ET_AL then
sep = comma and ', ' or ' & '
comma = true
end
end
authorstr = sep .. authors[i] .. authorstr
end
return authorstr
end
end
L.authorString = authorString
-- create the taxon authors string, including year, ex authors and authors of the basionym
local function createAllAuthorsStr(item, namequalifiers, year)
local authors = authorString(item, namequalifiers)
local authorsstr = ''
if authors or not year == '????' then
if code == NOMENCLATURE_ICNafp then
-- check for basionym
local basionymids = targetId(item, P_BASIONYM)
local basionymstr = ''
if next(basionymids) then
local basionym = mw.wikibase.getEntity('Q' .. next(basionymids))
local _,basionymnamequalifiers = targetStr(basionym, P_TAXON_NAME)
basionymstr = createAllAuthorsStr(basionym, basionymnamequalifiers)
if basionymstr ~= '' then
basionymstr = '(' .. basionymstr .. ') '
else
-- indicate missing basionym author
basionymstr = '(????) '
end
end
-- check ex-authors
local exauthors = authorString(nil, namequalifiers, P_EX_AUTHOR)
exauthorsstr = ''
mw.log(exauthors)
if exauthors then
exauthorsstr = exauthors .. ' ex '
end
authorsstr = basionymstr .. exauthorsstr .. authors
if year then
authorsstr = authorsstr .. ' (' .. year .. ')'
end
else
if year then
authorsstr = authors .. ', ' .. year
end
-- parentheses needed if instance of recombination
local recombination = false
for _,tid in pairs(qualifierTargetId(namequalifiers, P_INSTANCE_OF)) do
if tid == RECOMBINATION then
recombination = true
end
end
if recombination then
authorsstr = '(' .. authorsstr .. ')'
end
end
else
-- check for original combination
local basionymids = targetId(item, P_ORIGINAL_COMBINATION)
local basionymstr = ''
if next(basionymids) then
local basionym = mw.wikibase.getEntity('Q' .. next(basionymids))
local _, basionymnamequalifiers = targetStr(basionym, P_TAXON_NAME)
local pubyear = qualifierTargetValue(basionymnamequalifiers, P_TAXON_YEAR) or
targetValue(basionym, P_TAXON_YEAR)
-- access year in time representation "+1758-00-00T00:00:00Z"
local year = pubyear and string.sub(pubyear.time, 2, 5) or '????'
basionymstr = createAllAuthorsStr(basionym, basionymnamequalifiers, year)
if basionymstr ~= '' then
basionymstr = '(' .. basionymstr .. ') '
end
end
authorsstr = basionymstr
end
return authorsstr
end
L.createAllAuthorsStr = createAllAuthorsStr
-- show the stratigraphic range in which an extinct fossil existed
local function fossilParams(item, params)
local era1, era1references = next(targetId(item, P_ERA_START))
local era2, era2references = next(targetId(item, P_ERA_END))
local era1value = targetValue(item, P_START_TIME)
local era1time = era1value and era1value.time
local era2value = targetValue(item, P_END_TIME)
local era2time = era2value and era2value.time
if era1 and not (era1 == 'novalue') or era1time and not (era1time == "0") then
if era1 and not (era1 == 'novalue') then
params["era[1][label]"] = getLabel(era1)
params["era[1][id]"] = era1
end
if era1time and not (era1time == "0") then
local year = string.match(era1time, "^-*(%d-)-")
params["era[1][time]"] = year and tonumber(year) / 1000000
end
if era2 and not (era2 == 'novalue') then
params["era[2][id]"] = era2
params["era[2][label]"] = params["era[1][label]"]
if not (era1 == era2) then
params["era[2][label]"] = getLabel(era2)
end
end
if era2time and not (era2time == "0") then
local year = string.match(era2time, "^-*(%d-)-")
params["era[2][time]"] = year and tonumber(year) / 1000000
elseif era1time and not era2 and not (era2 == 'novalue') then
params["era[2][time]"] = "0"
end
-- merge references from era2 to era1, only show once
if era2 and not (era2 == 'novalue') and era2references and era2references.refids then
for a, b in pairs(era2references.refids) do
era1references.refids[a] = b
end
end
-- TODO: return data structure instead of pure str here
if era1references and era1references.refids then
params["era[references]"] = era1references.refids
end
end
end
L.fossilParams = fossilParams
-- returns html for the given refids set
-- parameters:
-- refids: list of integer ID to create a list of <ref>-references
local function references(refids)
mw.log('references support disabled. Import Cite module to make it work')
--
--local frame = mw.getCurrentFrame()
local refstr = ''
--if refids then
-- for id,_ in pairs(refids) do
-- local ref = Cite.citeitem(id, getLang()) or 'Error during creation of citation. Please report [[' .. id .. ']] at [[Module_talk:Cite]]'
-- mw.log('refstr for ', id, ref)
-- refstr = refstr .. frame:extensionTag('ref', ref, {name=id})
-- end
--end
return refstr
--
end
L.references = references
local function i18nByLatin(ranklatin, str, default)
local suc, format = pcall(i18n, str .. "-" .. ranklatin)
if not suc then
format = default
end
return format
end
L.i18nByLatin = i18nByLatin
local function formatScientificName(ranklatin, scientific, short)
local pf = "scientific-name"
if short then
pf = "short-" .. pf
end
scipattern = i18nByLatin(
ranklatin, pf .. "-pattern", i18n(pf .. "-pattern"))
scirepl = i18nByLatin(
ranklatin, pf .. "-repl", i18n(pf .. "-repl"))
scientific = string.gsub(scientific, scipattern, scirepl)
for scipattern, scirepl in pairs(
i18nByLatin(ranklatin, pf .. "-replaces", i18n(pf .. "-replaces"))) do
scientific = string.gsub(scientific, scipattern, scirepl)
end
return scientific
end
L.formatScientificName = formatScientificName
local function renderTableHead(text, color, extra_css)
local css = "text-align: center;"
if extra_css then
css = css .. " " .. extra_css
end
if color then
css = css .. " background-color: " .. color .. ";"
end
return mw.text.tag('tr', {}, mw.text.tag('th', {
colspan='2', style=css}, text))
end
L.renderTableHead = renderTableHead
local function renderTableRow(text, extra_css)
local css = "text-align: center;"
if extra_css then
css = css .. " " .. extra_css
end
return mw.text.tag('tr', {}, mw.text.tag('td',
{colspan='2', style=css}, text))
end
L.renderTableRow = renderTableRow
-- wiki-based projects urls
local function commons(text, color, extra_css)
local css = "text-align: center;"
local row
local species
local category
species = "[[File:Wikispecies-logo.svg|17px]]<br>[[species:" .. text .."|'''" .. getLabel(P_TAXON_NAME) .. "''']]"
category = "[[File:Commons-logo.svg|15px]]<br>[[c:category:" .. text .."|'''" .. getLabel(COMMONS_CATEGORY) .. "''']]"
if extra_css then
css = css .. " " .. extra_css
end
if color then
css = css .. " background-color: " .. color .. ";"
end
row = mw.text.tag('tr',{},
mw.text.tag('td', {colspan='2', style=css},
mw.text.tag('table', {width='100%'},
mw.text.tag('tr', {},
mw.text.tag('td', {}, species) ..
mw.text.tag('td', {}, category)))))
return row
end
L.commons = commons
local function renderFossilEra(params)
local eralink = {}
local refstr = references(params["era[references]"])
for i = 1, 2 do
local eraid = params[string.format("era[%d][id]", i)]
if eraid then
eralink[#eralink + 1] = getLink(eraid)
end
end
local separator = i18n("era-separator")
return renderTableHead(getLink(GEOLOGICAL_ERA) .. refstr, params.color) ..
renderTableRow(table.concat(eralink, separator))
end
L.renderFossilEra = renderFossilEra
local function renderIUCNStatus(params)
local r = {}
local refstr = references(params["iucn_status[references]"])
r[#r + 1] = renderTableHead(
getLink(RED_DATA_LIST, getLabel(P_IUCN_STATUS)) .. refstr, params.color)
r[#r + 1] = renderTableRow(
"[[File:" .. params["iucn_status[image]"] ..
"|lang=" .. getLang() ..
"|270px|" .. params["iucn_status[label]"] .. "]]")
return table.concat(r)
end
L.renderIUCNStatus = renderIUCNStatus
local function formatTaxon(
latin, qid, scientific, vernacular, is_subject, is_extinct)
local nameformat
scientific = scientific or i18n("no-scientific-name")
local scientificshort = scientific
if latin then
scientificshort = formatScientificName(latin, scientific, code ~= NOMENCLATURE_ICVCN)
scientific = formatScientificName(latin, scientific)
end
local nf = "item-format-parent"
if is_subject then
nf = "item-format-current"
end
if vernacular then
nameformat = i18n(nf .. "-with-vernacular-name")
else
nameformat = i18n(nf .. "-without-vernacular-name")
end
local link = qid
if _linkconfig == "sitelink" then
link = mw.wikibase.sitelink(qid) or "d:" .. qid
end
if is_extinct then
nameformat = i18n("extinct-mark") .. nameformat
end
return namedStringFormat{
nameformat, link=link, vernacular=vernacular,
scientific=scientific, scientificshort=scientificshort},
scientific
end
L.formatTaxon = formatTaxon
local function renderRank(i, params)
local row
local detailrows = {}
local pf = string.format("rank[%d]", i)
local ranklink = i18n("unknown-rank")
local rankid = params[pf .. "[id]"]
local ranklatin = params[pf .. "[latin]"]
local is_subject = params[pf .. "[is_subject]"]
local scientific = params[pf .. "[scientific]"]
local formatted = params[pf .. "[taxon]"]
if code == NOMENCLATURE_ICVCN and rankid == 22666877 then -- superdomain
return nil, detailrows
end
if code == NOMENCLATURE_ICVCN and ranklatin == "regnum" then
ranklink = getLink(VIRUS_CLASSIFICATION, i18n("virus-group-rank"), i18n("rank-format"), true, true)
local virusgenome = params["virus[genome]"]
if virusgenome then
local group = virusgroups[virusgenome] and virusgroups[virusgenome].group
local shortlabel = virusgroups[virusgenome] and virusgroups[virusgenome].shortlabel or ""
local qid = "Q" .. virusgenome
local link = (_linkconfig == "sitelink")
and (mw.wikibase.sitelink(qid) or "d:" .. qid)
or qid
local label = getLabel(virusgenome)
formatted = namedStringFormat{
i18n("virus-item-" .. (group and "with" or "without") .. "-group"),
group = group, link = link, shortlabel = shortlabel, label = label}
else
formatted = i18n("unknown-group")
end
else
if rankid then
local linkformat = i18nByLatin(
ranklatin, "rank-format", i18n("rank-format"))
ranklink = getLink(rankid, nil, linkformat, true, true)
end
if is_subject then
local refstr = references(params[pf .. "[references]"])
local authority = params[pf .. "[authority]"]
if code ~= NOMENCLATURE_ICVCN then
detailrows = {
renderTableHead(capitalize(
string.format(i18n("scientific-name-of-taxon"), getLabel(rankid)))
.. refstr, params.color
),
renderTableRow(scientific),
renderTableRow(authority, "font-variant: small-caps;")
}
elseif authority then
detailrows = {renderTableRow(authority, "font-variant: small-caps;")}
end
end
end
row = mw.text.tag(
'tr', {style="vertical-align: top;"},
mw.text.tag('td', {}, ranklink) ..
mw.text.tag('td', {}, formatted))
return row, detailrows
end
L.renderRank = renderRank
-- in case of more than one parent taxa or rank: choose target according to the
-- references selected by usereferences
local function chooseByRef(item, property)
local cand
local nextparent = {}
for id,refs in pairs(targetId(item, property)) do
-- some taxon, like Q2382443, the parent taxon is null
local novalue = id == "novalue"
-- try to find match from usetaxa
if not novalue and usetaxa["Q" .. id] then
table.insert(nextparent, {usetaxa["Q" .. id], id, refs})
end
-- or according to usereferences
if refs and refs.refids and type(refs.refids) ~= "boolean" then
for r, i in pairs(usereferences) do
if refs.refids[r] then
table.insert(nextparent, {i + usetaxa.size, id, refs})
end
end
end
if not novalue and not cand then -- if no item had references yet
cand = {nil, id, refs} -- use this
end
end
-- nextparent is not sorted, so sort it
table.sort(nextparent, function(a, b)
return a[1] < b[1]
end)
if next(nextparent) then
_, cand = next(nextparent)
end
if cand and cand[1] == nil and cand[3] and cand[3].refids then
for targetid, _ in pairs(cand[3].refids) do
usereferences.size = usereferences.size + 1
usereferences[targetid] = usereferences.size
end
end
if cand then
return cand[2], cand[3] or {}
else
return nil, {}
end
end
L.chooseByRef = chooseByRef
-- Find out if this taxon is extinct already
local function isExtinct(item)
-- check IUCN status
local statusid, _ = next(targetId(item, P_IUCN_STATUS))
if statusid == EXTINCT then
return true
end
-- check temporal range end
local eraend, _ = next(targetId(item, P_ERA_END))
if eraend and not eraend == 'novalue' then
return true
end
-- check end time
local endtime = targetValue(item, P_END_TIME)
if endtime then
return true
end
-- check instance of fossil taxon
if next(targetId(item, P_INSTANCE_OF)) == FOSSIL_TAXON then
return true
end
return false
end
L.isExtinct = isExtinct
-- Find out if the item is a monotypic taxon
local function isMonotypic(item)
return next(targetId(item, P_INSTANCE_OF)) == MONOTYPIC_TAXON
end
L.isMonotypic = isMonotypic
local function renderSynonyms(params)
local pf = string.format("rank[%d]", params["rank[size]"])
local ranklatin = params[pf .. "[latin]"]
local rows = ""
for i = 1, params["synonym[size]"] do
pf = string.format("synonym[%d]", i)
local synonym = namedStringFormat{i18n("item-format-synonym-render"),
link = params[pf .. "[link]"],
scientific = formatScientificName(ranklatin, params[pf .. "[name]"]),
author = params[pf .. "[author]"]}
rows = rows .. mw.text.tag('li', {}, synonym)
end
return rows
end
L.renderSynonyms = renderSynonyms
local function fetchDetails(qid, item, include_basionym_author, fetch_author)
item = item or mw.wikibase.getEntity(qid)
local name, namequalifiers, namereferences = targetStr(item, P_TAXON_NAME)
local link = qid
if _linkconfig == "sitelink" then
link = mw.wikibase.sitelink(qid) or "d:" .. qid
end
local authorsstr
if fetch_author then
local pubyear = qualifierTargetValue(namequalifiers, P_TAXON_YEAR) or
targetValue(item, P_TAXON_YEAR)
-- access year in time representation "+1758-00-00T00:00:00Z"
local year = pubyear and string.sub(pubyear.time, 2, 5) or '????'
-- basionym author can be suppressed by not providing an item to search in:
authorsstr = createAllAuthorsStr(include_basionym_author and item, namequalifiers, year)
end
return name, link, authorsstr, namereferences
end
L.fetchDetails = fetchDetails
local function getRank(item, id)
local rankid = id or chooseByRef(item, P_TAXON_RANK)
local ranklatin
if not rankid or rankid == "novalue" then
rankid = CLADE
end
if rankid then
ranklatin = getLabel(rankid, 'la')
ranklatin = ranklatin and mw.ustring.lower(ranklatin)
if rankid == ZOOSECTIO or rankid == ZOOSUBSECTIO then
ranklatin = 'zoo' .. ranklatin
end
end
return rankid, ranklatin
end
local function taxonParams(qid, item, params, fetch_detail, is_subject, is_parent_extinct, incertae_sedis_ranks)
local rankid
local ranklatin
local level = params["rank[size]"] + 1
local pf = string.format("rank[%d]", level)
local is_extinct = is_parent_extinct or isExtinct(item)
if #incertae_sedis_ranks > 0 then
local incertae_sedis_vernacular = getLabel(INCERTAE_SEDIS)
local raw_scientific = getLabel(INCERTAE_SEDIS, 'la')
local incertae_sedis_formatted = getLink(INCERTAE_SEDIS, nil, i18n("item-format-incertae-sedis"), true)
for i = #incertae_sedis_ranks, 1, -1 do
rankid, ranklatin = getRank(nil, incertae_sedis_ranks[i])
if not (hideranks[rankid] or hideranks[ranklatin]) then
params["rank[size]"] = level
params[pf .. "[id]"] = rankid
params[pf .. "[link]"] = "Q" .. INCERTAE_SEDIS
params[pf .. "[vernacular]"] = incertae_sedis_vernacular
params[pf .. "[raw_scientific]"] = raw_scientific
params[pf .. "[latin]"] = ranklatin
params[pf .. "[scientific]"] = raw_scientific
params[pf .. "[taxon]"] = incertae_sedis_formatted
level = level + 1
pf = string.format("rank[%d]", level)
end
end
end
local name, link, authorsstr, namereferences = fetchDetails(qid, item, true, fetch_detail)
params[pf .. "[references]"] = namereferences
params[pf .. "[authority]"] = authorsstr
local vernacular = vernacularName(item)
rankid, ranklatin = getRank(item)
if rankid == SUBGENUS and
string.match(name, "^%w+$") and
params[string.format("rank[%d][id]", level - 1)] == GENUS then
-- follow ICZN to prepend genus name in front of subgenus name
name = string.format("%s (%s)",
params[string.format("rank[%d][raw_scientific]", level - 1)],
capitalize(mw.ustring.lower(name)))
end
if (hideranks[rankid] or hideranks[ranklatin]) and not usetaxa[qid] then
-- interrupt since this rank has been hided from display
return params, is_extinct
end
name = name and capitalize(name)
local ranklatinformat = (code == NOMENCLATURE_ICVCN) and "virus" or ranklatin
local formatted, sciname = formatTaxon(
ranklatinformat, qid, name, vernacular, is_subject, is_extinct)
params["rank[size]"] = level
params[pf .. "[id]"] = rankid
params[pf .. "[link]"] = qid
params[pf .. "[is_monotypic]"] = isMonotypic(item)
params[pf .. "[vernacular]"] = vernacular
params[pf .. "[raw_scientific]"] = name
params[pf .. "[latin]"] = ranklatin
params[pf .. "[is_extinct]"] = is_extinct
params[pf .. "[scientific]"] = sciname
params[pf .. "[is_subject]"] = fetch_detail
params[pf .. "[taxon]"] = formatted
return params, is_extinct
end
L.taxonParams = taxonParams
-- performs the loop up the hierarchy using P_TAXON_PARENT
local function iterateRanks(
qid, count, fetch_detail, child_detailed, child_extinct, params)
local params = params or {["rank[size]"] = 0, ["synonym[size]"] = 0}
local item = mw.wikibase.getEntity(qid)
name = targetStr(item, P_TAXON_NAME)
if name == 'nil' then
return params, {}, item
end
if not code then
codeid = next(targetId(item, P_NOMENCLATURE_CODE))
if codeid and colors[codeid] then
code = codeid
end
end
if not subcode and colors[tonumber(string.match(qid, "^Q(%d+)"))] then
subcode = tonumber(string.match(qid, "^Q(%d+)"))
end
params["code"] = params["code"] or subcode or code
params["color"] = colors[params["code"]]
if not params["virus[genome]"] then
local genomeid, genomereferences = next(targetId(item, P_VIRUS_GENOME))
params["virus[genome]"] = genomeid
if genomereferences and genomereferences.refids then
params["virus[references]"] = genomereferences.refids
end
end
local nextid, refsquals = chooseByRef(item, P_TAXON_PARENT)
mw.log('nextid', nextid)
if visited[nextid] then -- loop detection
return params, {}, item
elseif nextid then
visited[nextid] = true
end
local is_subject = fetch_detail
-- Monotypic taxon can contain extinct taxa,
-- should not fetch detail in such circumstances
-- for example: [[Q7105303]]
local fetch_detail =
fetch_detail or
(child_detailed and not child_extinct and isMonotypic(item))
local is_extinct, is_parent_extinct
if nextid and (not code or count > 0) then
params, refs, _, is_parent_extinct = iterateRanks(
'Q' .. nextid, count - 1, false,
fetch_detail, isExtinct(item), params)
if refs then
for ref, _ in pairs(refs) do
refsquals.refids[ref] = true
end
end
end
if count > 0 then
incertae_sedis_ranks = qualifierTargetId(refsquals.qualifiers, P_INCERTAE_SEDIS)
params, is_extinct = taxonParams(
qid, item, params, fetch_detail, is_subject, is_parent_extinct, incertae_sedis_ranks)
end
return params, refsquals.refids, item, is_extinct
end
L.iterateRanks = iterateRanks
local function cladusPostfixes(n)
local plus = ""
for i = 1, n do
plus = plus .. "+"
end
return plus
end
-- use arguments from second table to override the first table
-- support classical {{taxobox}} parameters like "species", "unranked_ordo"
local function overrideParams(params, overrides)
overrides = overrides or {}
for key, val in pairs(overrides) do
params[key] = overrides[key] or params[key]
end
-- classical taxonomic rank params
local unranked = {}
for i = 1, params["rank[size]"] do
local pf = string.format("rank[%d]", i)
local latin = params[pf .. "[latin]"]
if latin == "clade" then
unranked[#unranked + 1] = i
else
local txarg = pf .. "[taxon]"
local atarg = pf .. "[authority]"
params[txarg] = latin and overrides[latin] or params[txarg]
params[atarg] = latin and overrides[latin .. "_authority"] or params[atarg]
for j = #unranked, 1, -1 do
local txarg = string.format("rank[%d][taxon]", unranked[j])
local atarg = string.format("rank[%d][authority]", unranked[j])
local argname = string.format("unranked_%s", latin)
if j == #unranked then
params[txarg] = overrides[argname] or overrides[argname .. "1"] or
overrides[latin .. "+"] or params[txarg]
params[atarg] = overrides[argname .. "_authority"] or
overrides[argname .. "1_authority"] or
overrides[latin .. "+_authority"] or params[atarg]
else
params[txarg] = overrides[string.format('%s%d', argname, #unranked - j + 1)]
or overrides[latin .. cladusPostfixes(#unranked - j + 1)]
or params[txarg]
params[atarg] = overrides[string.format('%s%d_authority', argname, #unranked - j + 1)]
or overrides[latin .. cladusPostfixes(#unranked - j + 1) .. "_authority"]
or params[atarg]
end
end
unranked = {}
end
if latin == "species" then
local scarg = pf .. "[scientific]"
params[scarg] = overrides["binomial"] or params[scarg]
elseif ({subspecies=true, varietas=true, forma=true})[latin] then
local scarg = pf .. "[scientific]"
params[scarg] = overrides["trinomial"] or params[scarg]
end
end
return params
end
L.overrideParams = overrideParams
local function getTypeTaxon(qid, item, params)
item = item or mw.wikibase.getEntity(qid)
params["image"] = params["image"] or targetStr(item, P_IMAGE)
params["audio"] = params["audio"] or targetStr(item, P_AUDIO)
params["taxon"] = params["taxon"] or targetStr(item, P_TAXON_NAME)
local id, refsquals = next(targetId(item, P_TAXONOMIC_TYPE))
if not id then return params, nil end
local typeitem = mw.wikibase.getEntity("Q" .. id)
local ranklatin
local rankid = next(targetId(typeitem, P_TAXON_RANK))
if rankid then
ranklatin = mw.wikibase.getEntity('Q' .. rankid):getLabel('la')
if ranklatin then
ranklatin = mw.ustring.lower(ranklatin)
end
end
if not (ranklatin == "genus" or ranklatin == "species") then return params, nil end
if ranklatin == "genus" then
params = getTypeTaxon("Q" .. id, typeitem, params)
end
local name, link, authorsstr, namereferences = fetchDetails("Q" .. id, nil, true, true)
if namereferences then
for ref, _ in pairs(namereferences) do
refsquals.refids[ref] = true
end
end
params["type[" .. ranklatin .. "][name]"] = name
params["type[" .. ranklatin .. "][link]"] = link
params["type[" .. ranklatin .. "][authority]"] = authorsstr
params["type[" .. ranklatin .. "][references]"] = refsquals and refsquals.refids
return params
end
L.getTypeTaxon = getTypeTaxon
local function iterateSynonyms(qid, item, params)
visited[tonumber(string.match(qid, "^Q(%d+)"))] = true
params["image"] = params["image"] or targetStr(item, P_IMAGE)
params["audio"] = params["audio"] or targetStr(item, P_AUDIO)
params["range_map"] = params["range_map"] or targetStr(item, P_SPREAD_MAP)
if not (params["era[1][label]"] or params["era[1][time]"]) then
fossilParams(item, params)
end
if not params["iucn_status[id]"] then
local statusid, statusreferences = next(targetId(item, P_IUCN_STATUS))
if statusid then
params["iucn_status[id]"] = statusid
params["iucn_status[references]"] = statusreferences.refids
local status = mw.wikibase.getEntity("Q" .. statusid)
params["iucn_status[image]"] = targetStr(status, P_IMAGE)
params["iucn_status[label]"] = getLabel(status)
end
end
if not params["rank[" .. params["rank[size]"] .. "][latin]"] then
local rankid, ranklatin = getRank(item)
params["rank[" .. params["rank[size]"] .. "][id]"] = rankid
params["rank[" .. params["rank[size]"] .. "][latin]"] = ranklatin
end
local synonyms = {}
-- forward synonym properties
for _, property in pairs({P_BASIONYM, P_SYNONYM, P_REPLACED_SYNONYM, P_ORIGINAL_COMBINATION}) do
for id, refsquals in pairs(targetId(item, property)) do
synonyms[id] = refsquals
end
end
-- inverse synonym properties
for _, property in pairs({P_SUBJECT_ROLE, P_INSTANCE_OF}) do
for id, refsquals in pairs(targetId(item, property)) do
if (id == PROTONYM or id == BASIONYM or id == SYNONYM_TAXON) and refsquals and refsquals.qualifiers then
for _, protoid in pairs(qualifierTargetId(refsquals.qualifiers, P_OF)) do
synonyms[protoid] = refsquals
end
end
end
end
for id, refsquals in pairs(synonyms) do
if id and not visited[id] then -- loop detection
local synonym_qid = "Q" .. id
local synonym_item = mw.wikibase.getEntity(synonym_qid)
local refs
params, refs = iterateSynonyms(synonym_qid, synonym_item, params)
local name, link, authorsstr, namereferences = fetchDetails(synonym_qid, synonym_item, true, true)
local level = params["synonym[size]"] + 1
local pf = string.format("synonym[%d]", level)
params[pf .. "[link]"] = link
params[pf .. "[name]"] = name or getLabel(synonym_item)
params[pf .. "[author]"] = authorsstr
params["synonym[size]"] = level
if refs then
for ref, _ in pairs(refs) do
refsquals.refids[ref] = true
end
end
if namereferences then
for ref, _ in pairs(namereferences) do
refsquals.refids[ref] = true
end
end
end
end
local ret_refs = refsquals and refsquals.refids
return params, ret_refs
end
L.iterateSynonyms = iterateSynonyms
-- fetch params should passed to taxobox for the given qid (e.g., qid=Q729412
-- for Heloderma) and count higher levels of the taxon hierarchy.
-- developers: use this method for tests in the debug console, e.g.,
-- p.localFunction("getTaxoboxParams")('Q729412', 5)
local function getTaxoboxParams(qid, count)
visited = {}
local params, references, item = iterateRanks(qid, count, true)
if params["rank[size]"] == 0 then
return {}
end
params["rank[references]"] = references
local scarg = string.format("rank[%d][scientific]", params["rank[size]"])
local vnarg = string.format("rank[%d][vernacular]", params["rank[size]"])
params["name"] = params[vnarg] or params[scarg]
params, synonym_references = iterateSynonyms(qid, item, params)
params["synonym[references]"] = synonym_references
params = getTypeTaxon(qid, item, params)
return params
end
L.getTaxoboxParams = getTaxoboxParams
local function callbackTaxobox(template, params, overrides, dryrun)
local content = {}
local frame = mw.getCurrentFrame()
params = overrideParams(params, overrides)
for key, val in pairs(params) do
if type(val) == "boolean" then
val = val and "yes" or "no"
elseif type(val) == "table" and
string.match(key, "%[references%]$") then
local refs = {}
for r, _ in pairs(val) do
table.insert(refs, r)
end
val = table.concat(refs, " ")
end
if dryrun then
content[#content + 1] = string.format(
"|%s = %s", key, val)
else
params[key] = val
end
end
if dryrun then
table.sort(content, function(a, b)
a = string.gsub(a, "%[(%d+)%]", function(i)
return "[" .. string.rep("0", 3 - #i) .. i .. "]"
end)
b = string.gsub(b, "%[(%d+)%]", function(i)
return "[" .. string.rep("0", 3 - #i) .. i .. "]"
end)
return a < b
end)
content = "{{" .. template .. "\n" ..
table.concat(content, "\n") .. "\n}}\n"
content = frame:callParserFunction("#tag", "nowiki", content)
return mw.text.tag("pre", {}, content)
else
return frame:expandTemplate{title=template, args=params}
end
end
L.callbackTaxobox = callbackTaxobox
-- creates the taxobox from giving params
local function renderTaxobox(params, overrides)
local content = {}
params = overrideParams(params, overrides)
local color = params.color
-- title
content[#content + 1] = renderTableHead(params.name, color)
-- image
if params.image then
content[#content + 1] = renderTableRow(
"[[File:" .. params.image .. "|lang=" .. getLang() .. "|270px|.]]")
end
-- fossil era
if params["era[1][id]"] then
content[#content + 1] = renderFossilEra(params)
end
-- ranks
if params["rank[size]"] > 0 then
local refstr = references(params["rank[references]"]) or ""
content[#content + 1] = renderTableHead(
capitalize(params["virus[genome]"]
and getLink(VIRUS_CLASSIFICATION, nil, nil, nil, true)
or getLink(SYSTEMATICS, nil, nil, nil, true))
.. refstr, color)
local taxondetails = {}
for i = 1, params["rank[size]"] do
local row, detailrows = renderRank(i, params)
content[#content + 1] = row
taxondetails[#taxondetails + 1] = table.concat(detailrows)
end
content[#content + 1] = table.concat(taxondetails)
end
-- synonyms
if params["synonym[size]"] > 0 then
local refstr = references(params["synonym[references]"])
content[#content + 1] = renderTableHead(
capitalize(getLink(SYNONYM_TAXON, nil, nil, nil, true)) .. refstr, color)
content[#content + 1] = mw.text.tag(
'tr', {colspan='2'}, mw.text.tag(
'td', {colspan='2'}, mw.text.tag(
'ul', {}, renderSynonyms(params))))
end
-- type taxons
for header, rank in pairs({[TYPE_GENUS] = "genus", [TYPE_SPECIES] = "species"}) do
if params["type[" .. rank .. "][name]"] then
local refstr = references(params["type[" .. rank .. "][references]"])
content[#content + 1] = renderTableHead(
capitalize(getLink(header, nil, nil, nil, true)) .. refstr, color)
local typetaxon = namedStringFormat{i18n("item-format-parent-without-vernacular-name"),
link = params["type[" .. rank .. "][link]"],
scientific = formatScientificName(rank, params["type[" .. rank .. "][name]"])}
content[#content + 1] = renderTableRow(typetaxon)
content[#content + 1] = renderTableRow(params["type[" .. rank .. "][authority]"], "font-variant: small-caps;")
end
end
-- subdivision
if params.subdivision and params.subdivision ~= "" then
content[#content + 1] = renderTableHead(i18n('subdivision-ranks'), color)
content[#content + 1] = mw.text.tag(
'tr', {colspan='2'}, mw.text.tag(
'td', {colspan='2', style=css}, params["subdivision"]))
end
-- range map
if params.range_map then
content[#content + 1] = renderTableHead(i18n('range-map'), color)
content[#content + 1] = renderTableRow(
"[[File:" .. params.range_map .. "|lang=" .. getLang() .. "|270px|.]]")
end
-- iucn status
if params["iucn_status[id]"] then
content[#content + 1] = renderIUCNStatus(params)
end
-- audio
if params.audio then
content[#content + 1] = renderTableHead(
getLink(P_AUDIO, nil, nil, nil, true), color)
content[#content + 1] =
renderTableRow("[[File:" .. params.audio .. "]]")
end
-- Wikispecies and Wikidata
if params.taxon then
content[#content + 1] =
commons(params.taxon, color)
end
return mw.text.tag('table', {
style = [[
width: 280px; border-width: 1px; float: right;
border-style: solid; background-color: #f9f9f9;
]]
}, table.concat(content))
end
L.renderTaxobox = renderTaxobox
if debug then
function p.debugParams(params)
mw.log("Start of logging params")
mw.log(string.rep("=", 20))
for k, v in pairs(params) do
mw.log(k, v)
end
mw.log("End of logging params")
end
function p.localFunction(name)
return L[name]
end
end
function p.taxobox(frame)
local config = parseConfig(frame.args)
local qid = frame.args.qid or mw.wikibase.getEntityIdForCurrentPage()
local params = getTaxoboxParams(qid, config.count)
return renderTaxobox(params, frame.args)
end
function p.callback(frame)
local config = parseConfig(frame.args)
local qid = frame.args.qid or mw.wikibase.getEntityIdForCurrentPage()
local template = frame.args.template or "Taxobox"
local params = getTaxoboxParams(qid, config.count)
return callbackTaxobox(template, params, frame.args, config.dryrun)
end
return p